Informatics
Our informatics scientists collaborate closely with other investigators and develop algorithms and software to disseminate research results to the broader community.
About Informatics
Contact
Informatics plays a key role in organizing, processing, analyzing, and interpreting the large quantity and variety of data generated in translational research. The Informatics (IFX) Core covers a wide array of expertise, including bioinformatics, multi-omics, cheminformatics, clinical informatics, data science, software engineering, UI/UX research and design, and project management. IFX Core team members collaborate extensively with informaticians embedded in other branches and cores within the Division of Preclinical Innovation (DPI) and with other colleagues across DPI to produce methodologies, resources and software for the translational research community.
IFX Core Mission
The mission of the IFX Core is to derive actionable insights from integrating translational research data and to accelerate the translation of findings into the clinic by:
- Identifying biological and chemical mechanisms that underlie diseases, including rare diseases, and their development, drug mechanisms of action and treatment response using novel or existing methods
- Improving use and interpretation of metabolomics and other omics datasets by developing new methods or enhancing the application of existing methods
- Producing open, comprehensive resources to accelerate translational research efforts, spanning ingredient/drug regulatory information, target annotations, disease annotations and molecular phenotypes
- Producing tools for the analysis and interpretation of complex high-throughput datasets
- Developing, optimizing and testing models to prioritize targets and therapeutic opportunities, and identifying repurposed drugs through collaborations with NCATS’ DPI branches
- Enhancing transparency and open research by adhering to user-centric and FAIR (Findable, Accessible, Interoperable, Reproducible) best practices
- Expanding the use and understanding of informatics in translational research through workshops, training and mentoring
What We Do
The work of the IFX Core falls into five main categories:
- Translational data analytics. Developing custom workflows and new methods to help explain complex, large-scale data sets that include multi-omic and clinical data; maintaining and deploying these tools internally and publicly.
- Rare disease translational research. Exploring many types of biomedical and clinical data to develop rare disease–based biomedical informatics applications; normalizing, harmonizing, integrating and representing rare disease–related data; retrieving and extracting rare disease–relevant information from free text; providing clinical decision support for rare diseases; applying computational drug repositioning to rare diseases.
- Standards, knowledge sources and software. Building, integrating, curating and publicly rendering resources to analyze various types of experimental and curated data sets.
- Metabolomics and multi-omics. Identifying presumed biomarkers and explaining disease processes through molecular profiling.
- Project management and outreach. Enabling the team to collaborate and deliver robust solutions and tools.
These five components are coordinated through our governance structure and efforts where one component informs efforts in another.
Related Research

Analytical Chemistry
Our analytical chemistry experts and state-of-the-art lab support early-stage chemical development.
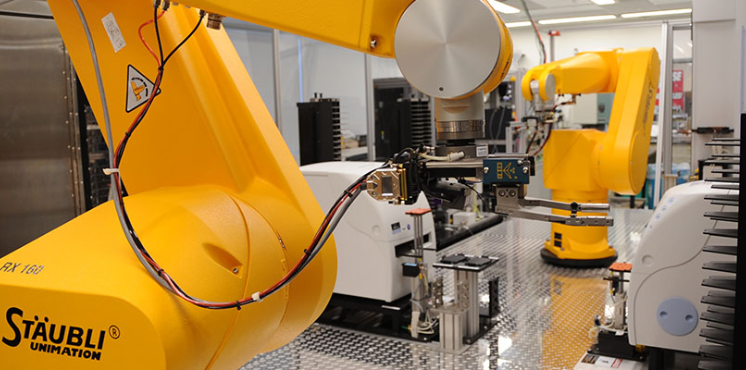

Automation
Our automation experts work closely with our scientists to support various research activities, including high-throughput screening and assay development and optimization.

Compound Management
Our compound management team uses sophisticated and automated techniques to support NCATS screening activities.