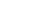Target Watch Sheds Light on Overlooked Proteins
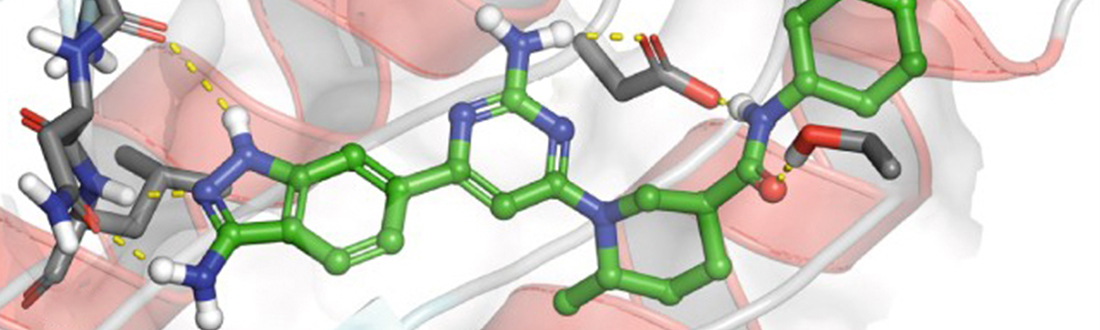
The Illuminating the Druggable Genome program is aimed at better understanding the potential of understudied proteins as targets for drugs. Some compounds and drugs that block the activity of proteins called kinases can have anticancer properties. Here, GSK2291363, a kinase inhibitor, shown with colored figures representing the different atoms and chemical bonds in the inhibitor’s chemical make-up, is docked with the protein kinase target, PIP5K1A, potentially blocking its activity. (Adapted by permission from Springer Nature Customer Service Centre GmbH: Springer Nature Nature Reviews Drug Discovery (PIP5K1A: a potential target for cancers with KRAS or TP53 mutations, Michael P. East, Tuomo Laitinen, Christopher R. M. Asquith, 2020))
July 31, 2020
Scientists estimate that approximately 3,000 genes in the human genome are “druggable,” meaning they make proteins that could be targets for existing or new drugs and compounds. Yet only a small percentage of proteins have been pursued in drug development. The Illuminating the Druggable Genome (IDG) program, supported by the NIH Common Fund and administered through NCATS and the National Institute of Diabetes and Digestive and Kidney Diseases, is aimed at shedding light on these understudied proteins and their potential use in developing treatments for diseases.
To draw attention to some of these “dark” proteins, IDG has partnered with the journal Nature Reviews Drug Discovery on a series of brief articles written by IDG scientists. The series, called Target Watch, provides a snapshot of understudied protein drug targets and their therapeutic potential.
“There are many parts of biology and medicine that we have yet to understand,” said Tudor Oprea, M.D., Ph.D., an IDG scientist at The University of New Mexico School of Medicine who oversees the series. The series is “a quick glimpse into the state of possibility.”
Each article focuses on a relatively underexplored area of drug target research. One such area is ion channels, protein passageways that allow sodium, potassium and other chemicals to flow in and out of cells and help the cells carry out their normal functions. Ion channels are promising therapeutic targets for treating many diseases. In a recent Target Watch article, investigators from the University of California, San Francisco, described their research on a class of ion channels for treating early-onset epilepsy, a disease that currently has limited treatment options. The team is developing tiny molecular probes that could alter channel activity.
Kinases, proteins that act as molecular switches, also represent another important area of interest for IDG scientists. These proteins turn different components on and off within pathways, chains of biological reactions that carry out activities in cells. Because defects in some pathways can lead to uncontrolled cell growth – a hallmark of cancer – many kinases are promising targets in cancer research. A team from The University of North Carolina at Chapel Hill School of Medicine and the University of Eastern Finland School of Pharmacy recently highlighted a type of kinase that affects fatty lipids in cells for its potential use in cancer therapy.
Target Watch also features research on novel protein targets for drugs that currently are on the market. In one article, Oprea and his colleagues described such targets, including nine that have implications for rare diseases. These targets include G-protein coupled receptors (GPCRs), a third major class of proteins under study by IDG scientists. GPCRs sit on the surface of cells and act like an inbox for chemical messages. Hundreds of types of GPCRs play many different roles in the body, including in sight, taste and smell and response to infection. They also are targets for many marketed drugs, including allergy and heart medications and antidepressants.
Target Watch provides a useful service to researchers. “In compiling this list, we’re highlighting new information on possible target proteins for the research community,” Oprea said. “Perhaps some scientists will consider studying a few themselves.”