NCATS-Supported Research Reduces Time to Diagnosis for Seriously Ill Children with Genetic Diseases
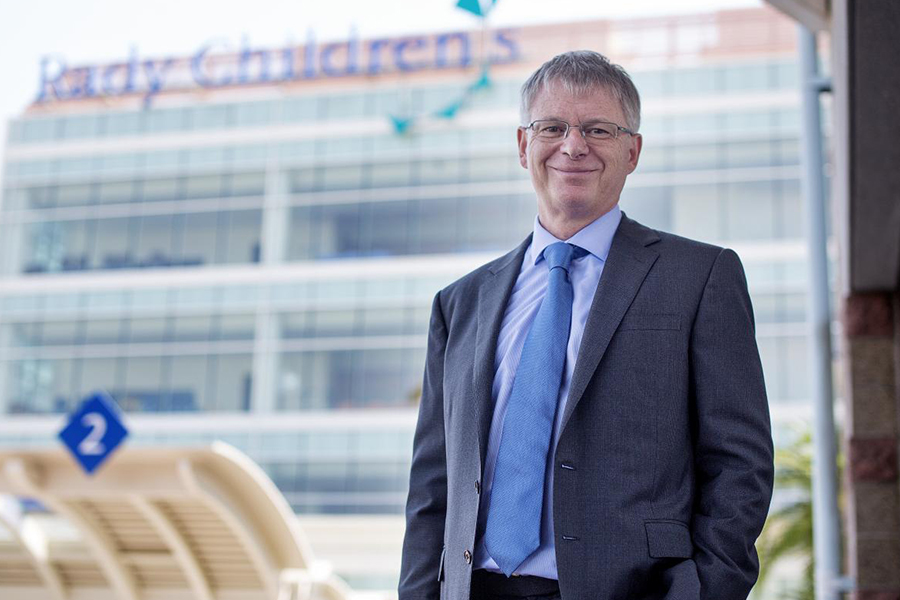

Dr. Stephen Kingsmore, President and CEO of Rady Children's Institute for Genomic Medicine
Credit: Rady Children's Institute for Genomic Medicine
March 11, 2022
Seriously ill children with genetic diseases, particularly infants in intensive care units for whom every hour and day is critical, might now be diagnosed and treated far more quickly than in the past.
In recent years, a technology called rapid whole-genome sequencing has been a successful first step to achieving faster diagnoses. The results of rapid whole-genome sequencing, however, must be interpreted by highly specialized individuals who are not always available when and where children need them. This has made it challenging to implement the technology at the point of care.
A group of researchers led by Stephen F. Kingsmore at the Rady Children’s Institute for Genomic Medicine has developed a new automated machine-learning approach for diagnosing these children more quickly—combining rapid whole-genome sequencing with automated phenotyping and interpretation. This approach helps achieve a diagnosis by comparing a patient’s genomic results with clinical information from electronic health records and elsewhere, reducing the need for labor-intensive manual analysis of genomic data. Automating this step significantly decreases the time to diagnosis and, consequently, the time to initiating appropriate treatment. The researchers described their results in a paper published in the April 24, 2019, issue of Science Translational Medicine. This new approach to diagnosing genetic diseases speeds answers to physicians caring for seriously ill children, ultimately leading to better outcomes.
This research was supported in part by an NCATS Clinical and Translational Science Award Program Collaborative Innovation Award. These awards are designed to support collaborative translational science innovations that benefit public health.