Our Staff
Our staff collaborate, innovate and speed research through a range of activities.
About Our Staff
Learn more about our staff and their work in support of the center’s mission.

Konstantinos Afratis, D.Phil.
Postdoctoral Visiting Fellow
Early Translation Branch
Division of Preclinical Innovation

Yong-Mo Ahn, Ph.D.
Research Scientist (Biology)
Early Translation Branch
Division of Preclinical Innovation




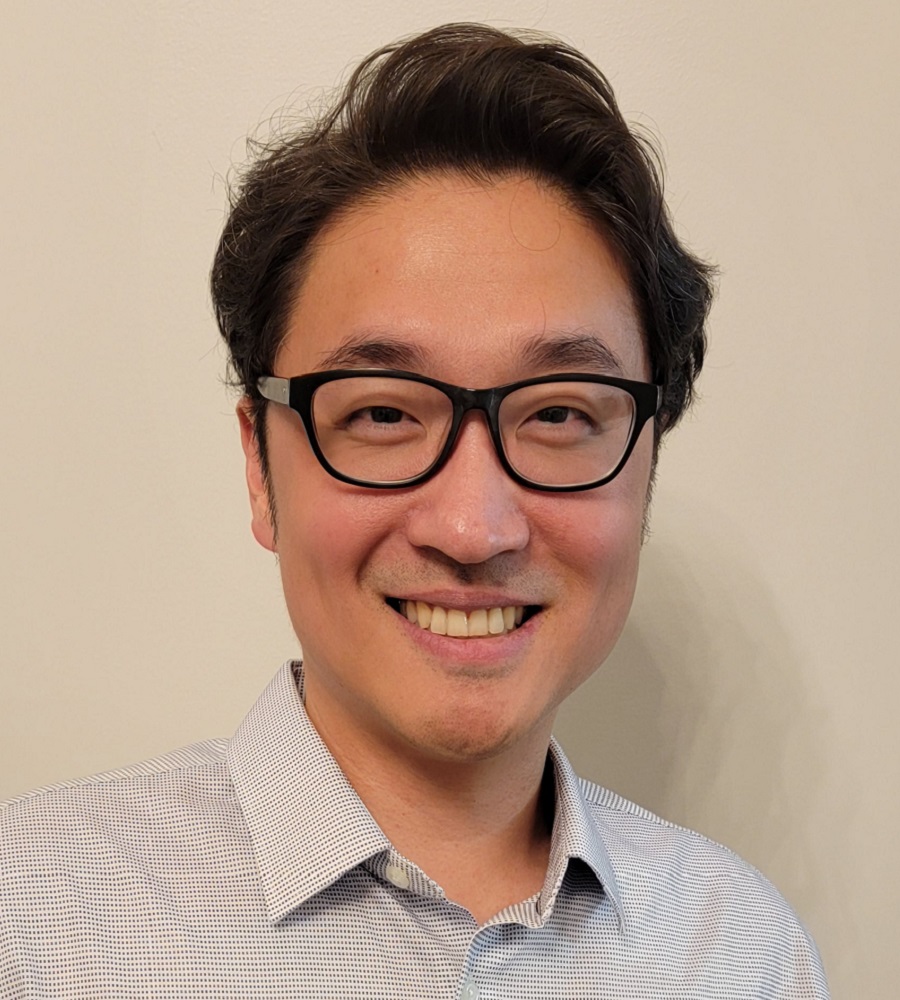
Dammika Nandanie Amugoda, M.S.
Curator/Research Scientist
Informatics
Division of Preclinical Innovation

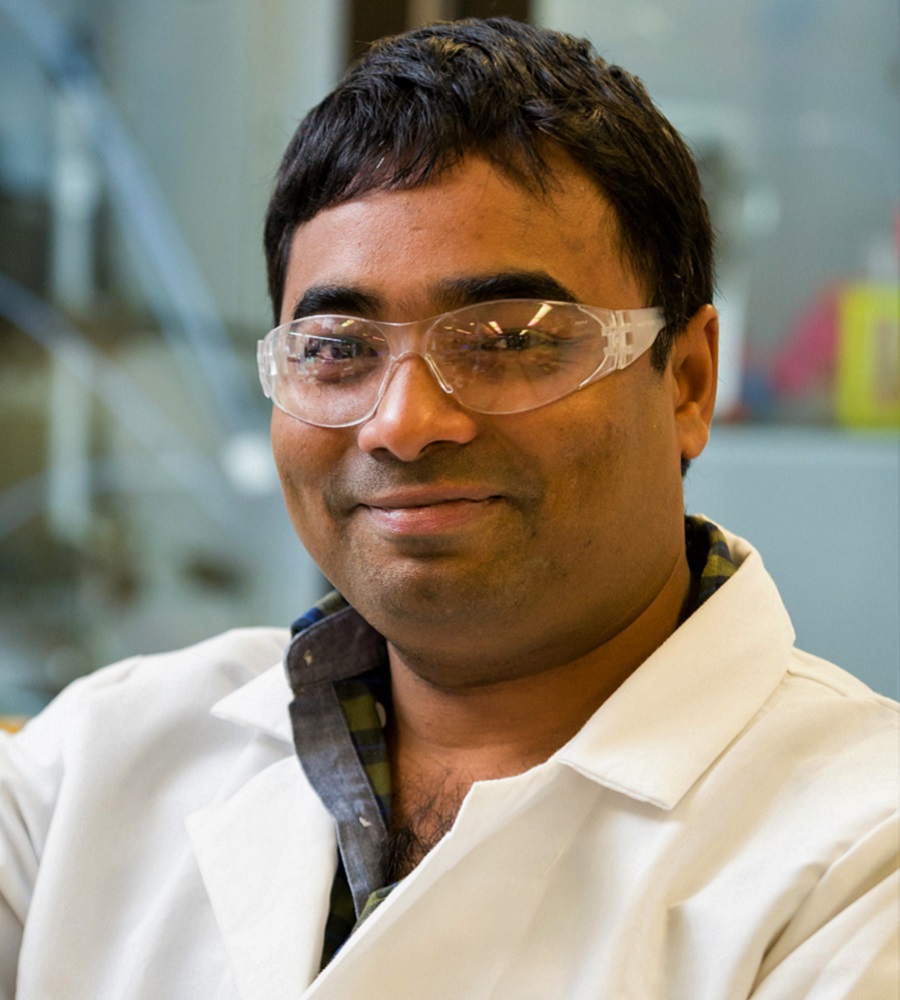
Guadalupe (Lupe) V. Aquino
Clinical Trials Specialist
Clinical Affairs Branch
Division of Clinical Innovation

Audie A. Atienza, Ph.D.
Program Officer
Clinical and Translational Science Awards Program Branch
Division of Clinical Innovation


Heather L. Baker
Lead Health Science Policy Analyst
Office of Program Evaluation, Analysis and Reporting
Division of Clinical Innovation





Jennifer M. Beierlein, Ph.D.
Health Science Policy Analyst
Policy Branch
Office of Policy, Communications and Education